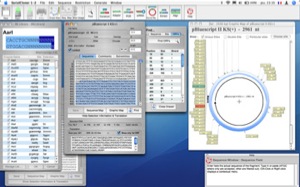
Inherit Features when copying/Pasting Protein sequences Non formatted copy of Protein sequence (Copy More… Menu) A rapid switch between a DNA and Protein Align window Inherit Features when reverse-translating a DNA sequence Scan for peptide Features in Protein Sequences A button/menu to choose particular sites to be displayed in Restriction maps. An option in the Restriction Map window to only translate nucleotides that are in uppercase An option in the Preferences to use the Codon code from Codon Usage tables as a Genetic code used for translation An option in the Preferences to select pK value tables to use for pI estimation MacOSX version now distributed as a signed application (Mountain Lion Gatekeeper compatibility) Estimates Isoelectric point of protein sequences (DNA and Protein Sequence Window) Insertion point position when editing a protein sequence Error in definition of PreScission site 
Slow down under certain Windows install due to autosaving Bug in "show info" toggle under Windows and Linux Detection of overlapping Primers in PCR a preference to limit or not the width of Seq & Prot windows (see the available contextual help window to get more details on new functions) It also allows to generate sense and antisense sequences to be obtained after bi-sulfite conversion. Serial Cloner 2.5 now allows to import codon frequency tables from the internet using a Web interface. It is also posible to send directly BLAST request at the NCBI and obtain the result inside a Web interface. You will also find a Restriction Enzyme library management interface, Additional tools, like a web browser for direct import of NCBI and EMBL entries, a virtual cutter to prepare restriction analysis or a silent restriction map generator to find how to introduce restriction sites without modifying the translated peptide are also provided. Features are now visible when aligning locally. Finally, Serial Cloner provides an interface to align two sequences using a local algorithm or the BLAST2Seq NCBI server. 
An additional interface allows easy Gateway(tm) cloning for both BP and LR reactions. Just select, blunt if you need, and click the Ligate button. Finally, you can assemble fragments, obtained by PCR, adaptor/shRNA synthesis or simply by graphically selecting fragments between restriction sites. shRNA constructions based on pre-defined scaffolds are also automated. *.zip, *.7z, *.gzip, *.gz, *.bz2, *.tar, *.PCR-based fragment or synthetic adaptors. Table 1: File Types Imported by SeqMan NGen for De Novo Assemblies and Templated Alignments (Special Workflows) Note: SeqMan NGen supports the export of data in. Below are the file types supported for import for both normal workflows and special workflows. SeqMan NGen is included within Lasergene Genomics. Image Files (*.png, *.jpg, *.gif, *.bmp 8)ġ File-Save will save back as original file type. These files are not supported through the menu system.ĩ Projects or files must contain protein sequence data.ġ0 Only the sequence, features and comments of the file will be read in.ġ3 Lasergene does not support ABI files without base calls.ġ4 Protean 3D supports version 3.2 of PDB files.ġ5 Available to support gap closure in SeqMan Ultra and for assembly in SeqMan NGen.ġ6 Supports *.txt and *.gp UniProt files only.įASTA Format (*.fas, *.fap, *.fna, *.fasta) All other files will be ignored.Ģ Multiple Lasergene DNA sequence files can be created by EditSeq, SeqMan Pro, and MegAlign.ģ FOF can not contain reference to another multiple sequence file format such as FastA, DNA* mseq, or Zip.ĥ Referred to collectively as “Lasergene Documents” in “Files of Type” or “Show pull-down menus”.ħ GenBank formatted features will be parsed.Ĩ Through “drag and drop” only. SeqMan NGen Transcriptome Files (*.transcriptome)ġ Only zip files containing ABI, SCF3, Lasergene protein or Lasergene DNA files. SeqMan NGen Assembly Projects (*.assembly) Protein Data Bank files (*.pdb, *.ent, *.pdb.gz, *.ent.gz, *.zip, *.txt) Imported File Types File FormatĤ54 Life Sciences output files (*.sff *.fas and *.qual)įASTA Format (*.fasta, *.fas, *.fap, *.nt, *.aa)įASTA Alignment with Gaps (*.fasta, *.fa, *.fas, *.fna, *.fap, *.nt, *.aa, *.frn, *.faa, *.ffn, *.mfa) Lasergene Molecular Biology and Lasergene Proteinīelow are the file formats supported for import and export by the applications within the Lasergene Molecular Biology and Lasergene Protein packages.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
